Introduction
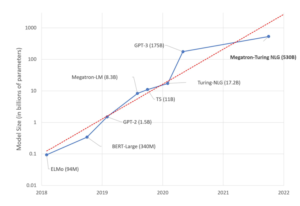
Often NLP datasets have large variations in their samples length. setting a maximum sequence length (max_seq_len) and pad shorter sequences with zeros is a common approach used with GPUs and CPUs.This approach is very inefficient as it results in many unrequired operations (multiply by zeros). The potential speed up by avoiding padding is the ratio of max_seq_len over average sequence length (avg_seq_len) in the NLP dataset. For MLPerf’s BERT dataset the potential speedup is roughly 2.
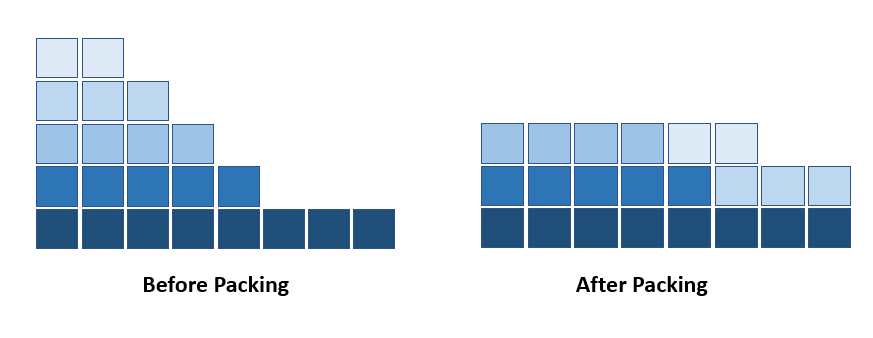
For Habana’s V1.1 MLPerf submission, we used a data packing technique [1], called Shortest-pack-first histogram-packing (SPFHP). Here instead of padding with zero, we pack several short sequences to one multi-sequence of size max_seq_len. Thus, we remove most of the padding, which can lead to up to x2 speedup in MLPerf BERT benchmark time-to-train (TTT). This packing technique can be applied on other types of datasets with high variability in samples length.
Please note that for each dataset with sequential data samples, the specific speedup with data packing is determined by the ratio of max_seq_len to average_seq_len in that particular NLP dataset. The larger the ratio, the higher the speedup.
Packing algorithm – Non-Negative Least-Square histogram:
(1) Go over the entire dataset and extract a histogram, H, of the sequence lengths
(2) Define a maximum number of sequences in a pack (we used 3) and find all possible combinations of sequence lengths that sum to max_seq_length. Those are all the possible strategies to pack.
(3) Create a matrix A of size max_seq_len×num_possible stratagies, where

(4) Solve iterativly:
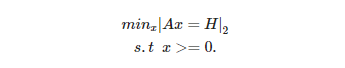
More technical details are in the code below and the paper [1]
Data format
When packing the data addtional fields should be added to the samples format to ensure mathematically equivlant operations. Below we detail the format changes.
Data format before packing:
- input_ids: size=512, list of token ids, where:
- 101 – start
- 102 – end
- 103 – mask
padded to 512 with zeros.
- input_mask: size=512, 111…111000…000, where the number of ones corresponds to the effective sample length, padded to 512 with zeros.
- segment_ids: size=512, 000…000111…111000…000, where first zeros correspond to the first sentence, ones to second sentence, and padded to 512 with zeros.
- masked_lm_positions: size=76, positions of masked tokens (103), padded to 76 with zeros.
- masked_lm_ids: size=76, token ids of masked tokens, padded to 76 with zeros.
- masked_lm_weights: size=76, 111…111000…000, number of ones equals to number of masked tokens.
- next_sentence_labels: size=1, 0 or 1, where 1 if sentence 2 is the next sentence of sentence 1.
Data format after packing:
- input_ids: size=512, list of token ids where:
- 101 – start
- 102 – end
- 103 – mask
padded to 512 with zeros.
Example of 2 packed samples: 101,…,102,…,102,101,…,102,…,102,0…0
where 101,…,102 first sentence, ,…,102 second sentence,
101,…,102 third sentence and ,…,102 forth sentence.
- input_mask: size=512, 111…111222…222000…000, where the number of ones corresponds to the first sample length, and the number of twos corresponds to the second sample length. (If there are 3 samples 1…12…23…30…0.)
- segment_ids: size=512, 000…000111…111000…000111…111000…000 where 000…000111…111 the first and the second samples, padded to 512 with zeros.
- positions: size=512, 0,1,2,3,…,\<length of first sample> – 1,0,1,2,3,…,\<length of second sample> – 1,0,0,…,0
- masked_lm_positions: size=79, positions of masked tokens (103), padded to 79 with zeros.
- masked_lm_ids: size=79, token ids of masked tokens, padded to 76 with zeros.
- masked_lm_weights: size=79, 111…111222…222000…000 where 111…111 corresponds to the first sample and 222…222 to the second. (If there are 3 samples 1…12…23…30…0.)
- next_sentence_positions: size=3, \<position of first sample>,\<position of second sample>,0 (corresponds to 101 positions)
- next_sentence_labels: size=3, 0 or 1, where 1 if sentence 2 is the next sentence of sentence 1.
- next_sentence_weights: size=3, 110 (If there are 3 samples 111)
Required Modifications to The Model
In addition to packing the data, minor changes should be applied to the model.
(1) Position embedding:
Position embedding is a matrix of the shape (512, 1024), where row i in this matrix corresponds to the token at position i in the sample. Before packing, all positions were in increasing order, and thus we could slice first n rows from the position embedding matrix. After packing, token positions are no longer increasing sequence. Instead (in the case of 3 packed samples) it is a stack of three increasing sequences (look at positions field). Consequently, instead of using a slice, we should select rows in the position embedding matrix according to positions field.
(2) Attention mask:
Before packing, an attention mask was used by attention layers to filter out all paddings. For example, if we had a sample with an effective length of n <= 512, in input_ids field we would have n token ids padded by zeros till 512. In this case, the attention mask would be looked like in the following image:
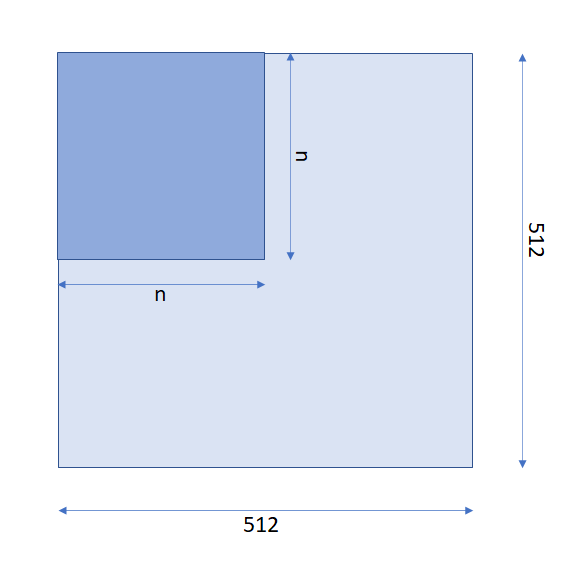
(512, 512) with the block of ones of shape (n, n) and all other elements are zeros.After packing, in addition to padding, we should filter out all cross-sample attentions, i.e., weight between tokens of two different samples And thus, for the case of 3 packed samples with the lengths
n, m and k the attention matrix should be looked like in the following image: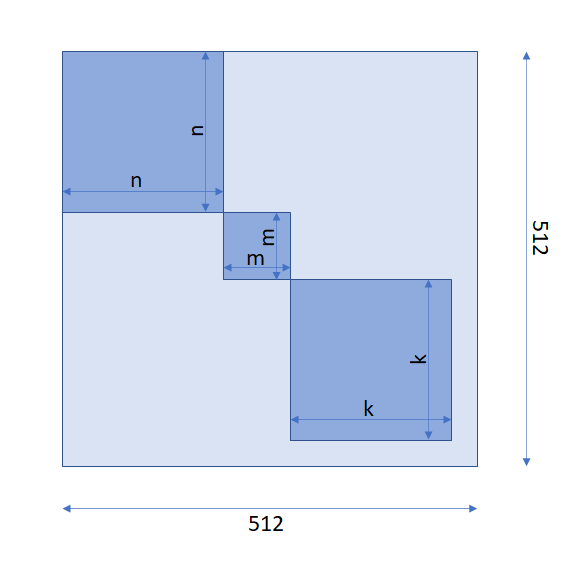
(512, 512) with blocks of ones of shapes (n, n), (m, m) and (k, k), and all other elements are zeros. We should use input_mask field to build such an attention mask matrix.(3) Pooler:
The output matrix of the model is a tensor of shape (512, 1024) (512 tokens and 1024 embedding dimension) i.e. 512 embedded tokens, where the first embedded token corresponds to 101 id. Before packing, only the first embedded token (101) was selected for the next sentence prediction task (the task where we should predict the sample’s following sentence), and thous we selected the first row in output matrix. After packing, we have up to 3 packed samples (where each multi-sample can be a pair of consecutive sentences), and thus, for each sample, we have also the position of its 101 token, which should be taken for the next sentence prediction task, therefore we should use next_sentence_positions field to select the correct embedded token matrix.
(4) Next sentence loss:
Before packing, there was only one pair of maybe consecutive sentences in each sample, thus we computed loss for each sample and averaged upon batch size. After packing, each multi-sample contains a different number of packed samples, from one to three, so we should average accross the effective number of samples in batch size. For that we should use next_sentence_weights field.
Examples
Next we dive into more detailed examples of the required model changes and the packing process.
Example 1: modifing the MLPerf’s BERT model
Specifically For MLPerf’s BERT model the following functions were modified:
1) In run_pretraining.py script input_fn_builder function: Since after packing we have different fields, we should change input_fn_builder.
Unpacked mode
:name_to_features = {
“input_ids”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“input_mask”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“segment_ids”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“masked_lm_positions”:
tf.io.FixedLenFeature([max_predictions_per_seq], tf.int64),
“masked_lm_ids”:
tf.io.FixedLenFeature([max_predictions_per_seq], tf.int64),
“masked_lm_weights”:
tf.io.FixedLenFeature([max_predictions_per_seq], tf.float32),
“next_sentence_labels”:
tf.io.FixedLenFeature([1], tf.int64),
}
Packed modePacked mode:
name_to_features = {
“input_ids”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“input_mask”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“segment_ids”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“positions”:
tf.io.FixedLenFeature([max_seq_length], tf.int64),
“masked_lm_positions”:
tf.io.FixedLenFeature([max_predictions_per_seq + 3], tf.int64),
“masked_lm_ids”:
tf.io.FixedLenFeature([max_predictions_per_seq + 3], tf.int64),
“masked_lm_weights”:
tf.io.FixedLenFeature([max_predictions_per_seq + 3], tf.float32),
“next_sentence_positions”:
tf.io.FixedLenFeature([3], tf.int64),
“next_sentence_labels”:
tf.io.FixedLenFeature([3], tf.int64),
“next_sentence_weights”:
tf.io.FixedLenFeature([3], tf.float32),
}
2) In run_pretraining.py script model_fn_builder function:
Unpacked modeUnpacked mode:
input_ids = features[“input_ids”]
input_mask = features[“input_mask”]
segment_ids = features[“segment_ids”]
masked_lm_positions = features[“masked_lm_positions”]
masked_lm_ids = features[“masked_lm_ids”]
masked_lm_weights = features[“masked_lm_weights”]
next_sentence_labels = features[“next_sentence_labels”]
Packed modePacked mode:
input_ids = features[“input_ids”]
input_mask = features[“input_mask”]
segment_ids = features[“segment_ids”]
masked_lm_positions = features[“masked_lm_positions”]
masked_lm_ids = features[“masked_lm_ids”]
masked_lm_weights = features[“masked_lm_weights”]
next_sentence_labels = features[“next_sentence_labels”]
positions = None
next_sentence_positions = None
next_sentence_weights = None
if FLAGS.enable_packed_data_mode and is_training: # only training should work in packed mode
positions = features[“positions”]
next_sentence_positions = features[“next_sentence_positions”]
next_sentence_weights = features[“next_sentence_weights”]
3) In modeling.py script embedding_postprocessor function: Added position embedding selection according to subsection in 1 Model changes part.
For packed model the following code was added:
flat_positions = tf.reshape(positions, [-1])
if use_one_hot_embeddings:
one_hot_positions = tf.one_hot(flat_positions, depth=seq_length)
position_embeddings = tf.matmul(one_hot_positions, position_embeddings)
else:
position_embeddings = tf.gather(position_embeddings, flat_positions)
position_embeddings = tf.reshape(position_embeddings, [batch_size, seq_length, width])
4) In modeling.py script create_attention_mask_from_input_mask function: Built block matrix according subsection 2 in Model changes part.
Unpacked mode:
to_shape = get_shape_list(to_mask, expected_rank=2) to_seq_length = to_shape[1]
to_mask = tf.cast(tf.reshape(to_mask, [batch_size, 1, to_seq_length]), tf.float32)
Packed mode:
to_mask = tf.one_hot(to_mask – 1, depth=3) to_mask = tf.matmul(to_mask, to_mask, transpose_b=True)
5) In modeling.py script BertModel.__init__ function pooler part: Selected first embedded token (101) for each packed sample according subsection 3 in Model changes part.
Unpacked mode:
selected_tokens = tf.squeeze(self.sequence_output[:, 0:1, :], axis=1)
Packed mode:
selected_tokens = gather_indexes(self.sequence_output, next_sentence_positions)
6) In run_pretraining.py script get_masked_lm_output function: Added float32 casting of masked_lm_weights field.
Packed mode:
label_weights = tf.cast(label_weights > 0, dtype=tf.float32)
7) In run_pretraining.py script get_next_sentence_output function: Applied change according to subsection 4 in Model changes part.
Unpacked mode:
loss = tf.reduce_mean(input_tensor=per_example_loss)
Packed mode:
weights = tf.reshape(weights, [-1]) numerator = tf.reduce_sum(input_tensor=weights * per_example_loss) denominator = tf.reduce_sum(input_tensor=weights) + 1e-5 loss = numerator / denominator
Example 2 : packing the TF records of a toy datatset
Next, we demonstrate how we addopted the code suggested by [1] and convert it to packed TF records. In the example below, we pack a toy dataset.
import os
import time
import glob
import struct
import random
import argparse
import numpy as np
import pandas as pd
from scipy import optimize
from itertools import repeat, chain
from functools import lru_cache, reduce
from collections import defaultdict, OrderedDict
from matplotlib import pyplot as plt
from concurrent.futures import ProcessPoolExecutor
import tensorflow as tfclass Args:
input_glob="/root/tensorflow_datasets/MLPerf_BERT_Wiki/unpacked_toy_data/"
output_dir="/root/tensorflow_datasets/MLPerf_BERT_Wiki/packed_toy_data/"
random_seed=12345
max_files=6 #default 100
duplication_factor=1
max_sequence_length=512
max_predictions_per_sequence=76
max_sequences_per_pack=3
args=Args()@lru_cache(maxsize=None)
def packing_strategies(start, previous, target, depth):
gap = target - start
# The collection of possible strategies given the
# starting sum, the target sum, and the available depth
# strategy search is limited to increments greater or equal to previous
strategies = []
# Complete the packing with exactly 1 number
if depth == 1:
if gap >= previous:
strategies.append([gap])
# Complete the sample in "depth" steps, recursively
else:
for new in range(previous, gap + 1):
new_gap = target - start - new
if new_gap == 0:
strategies.append([new])
else:
options = packing_strategies(start + new, new, target, depth - 1)
for option in options:
if len(option) > 0:
strategies.append([new] + option)
return strategiesdef get_packing_recipe(sequence_lengths, max_sequence_length, max_sequences_per_pack=3):
# Histogram of sequence lengths
histogram, bins = np.histogram(sequence_lengths, bins=np.arange(1, max_sequence_length + 2))
print("Begin packing pass".center(80, "_"))
print(f"Unpacked mean sequence length: {sequence_lengths.mean():3.2f}")
# Make sure all strategies are recipes to pack to the correct sequence length
strategy_set = packing_strategies(0, 1, max_sequence_length, max_sequences_per_pack)
for strategy in strategy_set:
assert(sum(strategy) == max_sequence_length)
num_strategies = len(strategy_set)
print(f"Found {num_strategies} unique packing strategies.")
# Solve the packing equation A@mixture = histogram
A = np.zeros((max_sequence_length, num_strategies), dtype=np.int32)
for i in range(num_strategies):
strategy = strategy_set[i]
for seq_len in strategy:
A[seq_len - 1, i] += 1
# short sequences are inexpensive to add, so should have low residual weights
# to exactly minimize padding use w0 = np.arange(1, max_sequence_length + 1)
# in practice the difference is negligible, but this converges faster
padding_cutoff = 8
w0 = np.ones([max_sequence_length])
# w0 = np.linspace(1, max_sequence_length+1, max_sequence_length)/max_sequence_length # padding minimization weight
w0[:padding_cutoff] = padding_cutoff / (2 * max_sequence_length)
w0 = np.sqrt(w0)
# Starting values for the padding and the mixture
padding = np.zeros([max_sequence_length], dtype=np.int32)
mixture = np.zeros([num_strategies], dtype=np.int32)
b = histogram + padding
# Pack sequences as best as possible, then increase padding accordingly and repeat
for i in range(0, 20):
print(f"\nIteration: {i}: sequences still to pack: ", b.sum())
start = time.time()
partial_mixture, rnorm = optimize.nnls(np.expand_dims(w0, -1) * A, w0 * b)
print(f"Solving nnls took {time.time() - start:3.2f} seconds.")
print(f"Residual norm: {rnorm:3.5e}")
# Update mixture (round the floating point solution to integers)
partial_mixture = np.where(partial_mixture < 2, np.rint(partial_mixture), np.floor(partial_mixture))
# If partial mixture is empty (due to rounding) we follow the gradient
# this usually happens when the number of examples is small i.e. ~100
if partial_mixture.max() == 0:
grad = A.T @ (b * np.arange(1, max_sequence_length + 1))
k = int(b.sum() // 2) + 1
topk = np.argsort(-grad)[:k]
partial_mixture[topk] += 1
# Update mixture
mixture = mixture + partial_mixture
# Compute the residuals
residual = b - A @ partial_mixture
print(f"Max residual: {abs(residual).max()}")
print(f"Residual on first 8 categories: {np.around(residual[:8], 4)}")
print(f"Residual on last 8 categories: {np.around(residual[-8:], 4)}")
# Add padding based on deficit (negative residual)
partial_padding = np.where(residual < 0, -residual, 0)
print(f"Added {(partial_padding*np.arange(1,max_sequence_length+1)).sum():3.2e} tokens of padding.")
padding = padding + partial_padding
# Update the rhs vector (remaining surplus sequences)
b = histogram + padding - A @ mixture
assert np.all(b >= 0), b
# Done iterating
if b.sum() < 100:
break
# Make sure there is no remainder
unpacked_seqlen = np.arange(1, args.max_sequence_length + 1)[b > 0]
# Update the mixture to also covered the unpacked sequences
for l in unpacked_seqlen:
# Get the depth 1 strategy
strategy = sorted([l, args.max_sequence_length - l])
strategy_index = strategy_set.index(strategy)
mixture[strategy_index] += b[l-1]
b = histogram - A @ mixture
padding = np.where(b < 0, -b, 0)
b = histogram + padding - A @ mixture
assert b.sum() == 0
# Analyze result
print("Done solving for packing order".center(80, "_"))
num_padding_tokens = (np.arange(1, max_sequence_length + 1) * padding).sum()
num_padding_tokens_original = (max_sequence_length - sequence_lengths).sum()
print(f"Number of sequences dropped: {b.sum()}")
print(f"Number of strategies utilized: {np.count_nonzero(mixture)}")
new_number_of_samples = int(mixture.sum())
compression = 1 - new_number_of_samples / len(sequence_lengths)
print(f"New number of samples: {new_number_of_samples:3.2f}, original {len(sequence_lengths)}. A compression ratio of {compression:3.3f}")
print(f"The expected speed-up from packing: {1/(1-compression):3.3f}")
upper_bound = 1.0 / (1 - ((1 - sequence_lengths / max_sequence_length).mean()))
print(f"Theoretical upper bound on speed-up: {upper_bound:3.3f}")
avg_sequences_per_sample = ((A.sum(0) * mixture).sum() - padding.sum()) / new_number_of_samples
print(f"Average sequences/sample {avg_sequences_per_sample:3.5f}")
print(f"Added {num_padding_tokens:3.2e} padding tokens. Original dataset used {num_padding_tokens_original:3.2e} padding tokens")
efficiency = (new_number_of_samples*max_sequence_length - num_padding_tokens)/(new_number_of_samples*max_sequence_length)
print(f"Packing efficiency (fraction of real tokens): {efficiency:3.4f}")
print(f"Top 8 strategies")
topK = np.argsort(-mixture)[:8]
for i in topK:
print(f"Strategy {strategy_set[i]} which is used {int(mixture[i])} times")
print("".center(80, "_"))
# Figure out the slicing that each strategy should use
slicing = np.zeros_like(A)
slicing[:, 1:] = np.cumsum(A * mixture, axis=1)[:, :-1]
slicing = slicing.T
mixture = mixture.astype(np.int64)
return strategy_set, mixture, padding, slicingdef slice_examples(examples_by_length, slicing, strategy_set, repeat_counts):
# Divide the work, firstly between the strategies and then into chunks of 50k
slices = []
strategies = []
part_idx = []
for strategy, slice_offsets, repeat_count in zip(strategy_set, slicing, repeat_counts):
if repeat_count == 0:
continue
# Slice out the sequences allocated to this strategy in increments of 50k
num_parts = repeat_count // 50000
num_parts = num_parts + int(repeat_count != num_parts * 50000)
subcounts = (min(50000, repeat_count - 50000 * (i - 1)) for i in range(1, num_parts + 1))
for part_id, part_count in enumerate(subcounts):
examples = []
for k, seq_len in enumerate(strategy):
slice_start = int(slice_offsets[seq_len - 1])
slice_end = slice_start + int(part_count)
slice_offsets[seq_len - 1] = slice_end
examples.append(examples_by_length[seq_len][slice_start:slice_end])
#import pdb; pdb.set_trace()
slices.append(examples)
strategies.append(strategy)
part_idx.append(part_id)
return slices, strategies, part_idxdef parallel_pack_according_to_strategy(args, part_idx, strategy, examples):
# Pack the sequences according to the strategy and write them to disk
base_filename = os.path.join(args.output_dir, "strategy_" + "_".join(map(str, strategy)))
filename = base_filename + f"_part_{part_idx}"
writer = tf.compat.v1.python_io.TFRecordWriter(filename)
for i, multi_sequence in enumerate(zip(*examples)):
features = create_multi_sequence_example(multi_sequence, args.max_predictions_per_sequence,
args.max_sequence_length, args.max_sequences_per_pack)
# Write to file
tf_example = tf.train.Example(features=tf.train.Features(feature=features))
writer.write(tf_example.SerializeToString())
writer.close()def create_multi_sequence_example(multi_sequence, max_predictions_per_sequence, max_sequence_length, max_sequences_per_pack):
# SEQ
packed_input_ids = np.zeros(max_sequence_length, dtype=np.int32)
packed_input_mask = np.zeros(max_sequence_length, dtype=np.int32)
packed_segment_ids = np.zeros(max_sequence_length, dtype=np.int32)
packed_positions = np.zeros(max_sequence_length, dtype=np.int32)
# MLM
# we are packing up to max_sequences_per_pack, each with a certain percentage of masked tokens
# in case that percentege is rounded up for all sequences in the pack, need to add an extra token for
# each sequence in the pack
packed_masked_lm_positions = np.zeros(max_predictions_per_sequence + max_sequences_per_pack, dtype=np.int32)
packed_masked_lm_ids = np.zeros(max_predictions_per_sequence + max_sequences_per_pack, dtype=np.int32)
packed_masked_lm_weights = np.zeros(max_predictions_per_sequence + max_sequences_per_pack, dtype=np.int32)
# NSP
packed_next_sentence_positions = np.zeros(max_sequences_per_pack, dtype=np.int32)
packed_next_sentence_labels = np.zeros(max_sequences_per_pack, dtype=np.int32)
packed_next_sentence_weights = np.zeros(max_sequences_per_pack, dtype=np.int32)
offset = 0
mlm_offset = 0
sequence_index = 1 # used in the input mask
for sequence in multi_sequence:
# Padding sequences are donoted with None
if sequence is not None:
example = tf.train.Example()
example.ParseFromString(sequence.numpy())
input_ids = np.array(example.features.feature['input_ids'].int64_list.value)
input_mask = np.array(example.features.feature['input_mask'].int64_list.value)
segment_ids = np.array(example.features.feature['segment_ids'].int64_list.value)
masked_lm_positions = np.array(example.features.feature['masked_lm_positions'].int64_list.value)
masked_lm_ids = np.array(example.features.feature['masked_lm_ids'].int64_list.value)
masked_lm_weights = np.array(example.features.feature['masked_lm_weights'].float_list.value)
next_sentence_labels = np.array(example.features.feature['next_sentence_labels'].int64_list.value)
#input_ids, input_mask, segment_ids, masked_lm_positions, masked_lm_ids, masked_lm_weights, next_sentence_labels = sequence
seq_len = input_mask.sum()
# SEQ
packed_input_ids[offset:offset + seq_len] = input_ids[:seq_len]
packed_input_mask[offset:offset + seq_len] = sequence_index
packed_segment_ids[offset:offset + seq_len] = segment_ids[:seq_len]
packed_positions[offset:offset + seq_len] = np.arange(0, seq_len)
# MLM
mlm_len = int(masked_lm_weights.sum())
assert mlm_offset + mlm_len < max_predictions_per_sequence + max_sequences_per_pack, "Too many LM predictions per sequences"
max_mlm = mlm_offset + mlm_len
#import pdb; pdb.set_trace()
packed_masked_lm_positions[mlm_offset:max_mlm] = offset + masked_lm_positions[:mlm_len]
packed_masked_lm_ids[mlm_offset:max_mlm] = masked_lm_ids[:mlm_len]
packed_masked_lm_weights[mlm_offset:max_mlm] = sequence_index
#import pdb; pdb.set_trace()
# NSP
packed_next_sentence_positions[sequence_index - 1] = offset
packed_next_sentence_labels[sequence_index - 1] = next_sentence_labels
packed_next_sentence_weights[sequence_index - 1] = 1
# Update offsets
sequence_index += 1
offset += seq_len
mlm_offset = max_mlm
#import pdb; pdb.set_trace()
# Pack into tfrecord format:
features = OrderedDict()
features["input_ids"] = create_int_feature(packed_input_ids)
features["input_mask"] = create_int_feature(packed_input_mask)
features["segment_ids"] = create_int_feature(packed_segment_ids)
features["positions"] = create_int_feature(packed_positions)
features["masked_lm_positions"] = create_int_feature(packed_masked_lm_positions)
features["masked_lm_ids"] = create_int_feature(packed_masked_lm_ids)
features["masked_lm_weights"] = create_float_feature(packed_masked_lm_weights)
features["next_sentence_positions"] = create_int_feature(packed_next_sentence_positions)
features["next_sentence_labels"] = create_int_feature(packed_next_sentence_labels)
features["next_sentence_weights"] = create_float_feature(packed_next_sentence_weights)
return features def create_int_feature(values):
feature = tf.train.Feature(int64_list=tf.train.Int64List(value=list(values)))
return featuredef create_float_feature(values):
feature = tf.train.Feature(float_list=tf.train.FloatList(value=list(values)))
return feature#path="/root/tensorflow_datasets/MLPerf_BERT_Wiki/unpacked_toy_data"#out="/root/tensorflow_datasets/MLPerf_BERT_Wiki/packed_toy_data"def main():
random.seed(args.random_seed)
# Put examples into bins depending on their sequence lengths and extract the sequence length
sequence_lengths = []
examples_by_length = defaultdict(list)
print("Looping through dataset to collect sequence length information...")
for filename in os.listdir(args.input_glob):
for record in tf.data.TFRecordDataset(args.input_glob+filename):
example = tf.train.Example()
example.ParseFromString(record.numpy())
im_length = sum(example.features.feature['input_mask'].int64_list.value)
examples_by_length[im_length].append(record)
sequence_lengths.append(im_length)
sequence_lengths = np.array(sequence_lengths)
# Pass the array of sequence lengths to the packing algorithm
strategy_set, mixture, padding, slicing = get_packing_recipe(sequence_lengths, args.max_sequence_length, args.max_sequences_per_pack)
# Add the calculated padding
for i in range(1, args.max_sequence_length + 1):
examples_by_length[i].extend([None] * int(padding[i - 1]))
# Shuffle the data
for key in examples_by_length:
random.shuffle(examples_by_length[key])
# Pack and store the data
print(f"\nPacking and writing packed dataset to {args.output_dir}.")
# Slice the data into chunks of max 50k packed examples
example_slices, strategies, part_idx = slice_examples(examples_by_length, slicing, strategy_set, mixture)
print(f"Splitting work into {len(part_idx)} parts.")
start = time.time()
#For debug uses
#for i in range(len(part_idx)):
# parallel_pack_according_to_strategy(args, part_idx[i], strategies[i], example_slices[i])
#import pdb; pdb.set_trace()
with ProcessPoolExecutor(16) as executor:
work = repeat(args), part_idx, strategies, example_slices
for partial_result in executor.map(parallel_pack_according_to_strategy, *work):
pass
print(f"\nDone. Took: {time.time() - start:3.2f} seconds to pack and write dataset.")main()Looping through dataset to collect sequence length information...
_______________________________Begin packing pass_______________________________
Unpacked mean sequence length: 392.13
Found 22102 unique packing strategies.
Iteration: 0: sequences still to pack: 1157748
Solving nnls took 22.57 seconds.
Residual norm: 5.50443e+02
Max residual: 3011.0
Residual on first 8 categories: [ -570. -755. -1131. -1507. -1883. -2259. -2634. -3011.]
Residual on last 8 categories: [21. 18. 15. 12. 9. 6. 5. 0.]
Added 9.01e+04 tokens of padding.
Iteration: 1: sequences still to pack: 1442.0
Solving nnls took 16.20 seconds.
Residual norm: 9.86222e+01
Max residual: 207.0
Residual on first 8 categories: [ -25. -51. -77. -102. -128. -154. -180. -207.]
Residual on last 8 categories: [2. 2. 2. 1. 1. 1. 1. 0.]
Added 1.22e+04 tokens of padding.
Iteration: 2: sequences still to pack: 263.0
Solving nnls took 14.64 seconds.
Residual norm: 3.38750e+01
Max residual: 24.0
Residual on first 8 categories: [ -4. -7. -10. -14. -16. -18. -23. -24.]
Residual on last 8 categories: [0. 0. 0. 0. 0. 0. 0. 0.]
Added 2.77e+03 tokens of padding.
Iteration: 3: sequences still to pack: 101.0
Solving nnls took 14.08 seconds.
Residual norm: 1.41189e+01
Max residual: 10.0
Residual on first 8 categories: [ 0. -3. -3. -4. -6. -6. -9. -10.]
Residual on last 8 categories: [0. 0. 0. 0. 0. 0. 0. 0.]
Added 9.33e+02 tokens of padding.
_________________________Done solving for packing order_________________________
Number of sequences dropped: 0.0
Number of strategies utilized: 692
New number of samples: 886919.00, original 1157748. A compression ratio of 0.234
The expected speed-up from packing: 1.305
Theoretical upper bound on speed-up: 1.306
Average sequences/sample 1.30536
Added 1.13e+05 padding tokens. Original dataset used 1.39e+08 padding tokens
Packing efficiency (fraction of real tokens): 0.9998
Top 8 strategies
Strategy [512] which is used 699578 times
Strategy [164, 169, 179] which is used 787 times
Strategy [67, 216, 229] which is used 771 times
Strategy [74, 201, 237] which is used 749 times
Strategy [77, 199, 236] which is used 740 times
Strategy [215, 297] which is used 733 times
Strategy [243, 269] which is used 710 times
Strategy [59, 196, 257] which is used 705 times
________________________________________________________________________________
Packing and writing packed dataset to /root/tensorflow_datasets/MLPerf_BERT_Wiki/packed_toy_data/.
Splitting work into 705 parts.
Done. Took: 89.37 seconds to pack and write dataset.Code language: JavaScript (javascript)Reference
Copyright (c) 2021 Habana Labs, Ltd. an Intel Company.
All rights reserved.
Licensed under the Apache License, Version 2.0 (the “License”);
you may not use this file except in compliance with the License. You may obtain a copy of the License at https://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an “AS IS” BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.